During vertebrate embryogenesis, fetal hematopoietic stem and progenitor cells (HSPCs) exhibit expansion and differentiation properties in a supportive hematopoietic niche (1, 2). Clinically, in vitro HSPC expansion is a feasible approach to obtain sufficient transplantable HSPCs but remains technically challenging (3, 4). Thus, decoding the complex regulatory mechanism of HSPC expansion within the hematopoietic organ is essential.
The caudal hematopoietic tissue (CHT) is characterized as a hematopoietic organ for fetal HSPC expansion in zebrafish (5). CHT is a highly vascularized niche, and the rapidly increased vascular endothelial cells (ECs) form a complex structure composing the dorsal artery and honeycomb-like venous network (6, 7). Therefore, the maintenance of CHT function relies on an orderly and precise process of interaction among all cell types inside (8). However, it remains to be resolved how global niche components orchestrate HSPC development.
Previously, Prof. Feng Liu’s group from the Institute of Zoology of the Chinese Academy of Sciences, has revealed the spatiotemporal characteristics of HSPC expansion in the CHT region using bulk RNA-sequencing (RNA-seq) and geographical position sequencing, however, our understanding of molecular and cellular dynamics in CHT hematopoiesis at single-cell resolution remains elusive.
In a recent study published in PNAS, Prof. Feng Liu 's group established the first developmental single-cell transcriptomic atlases of HSPC expansion in zebrafish and provides an essential resource for understanding regulatory mechanisms of HSPC expansion and differentiation in the niche.
The researchers first used single-cell RNA-seq (scRNA-seq) to map the transcriptional profiles of 28,777 cells, which were sorted from the CHT spanning three successive developmental stages. With these profiles, they resolved fetal HSPC heterogeneity, manifested as lineage priming and metabolic gene signatures. Furthermore, they investigated the regulatory mechanism of CHT niche components for HSPC development, with a focus on the transcription factors and ligand–receptor networks involved in HSPC expansion. Importantly, they identified an EC-specific G protein–coupled receptor 182, followed by in vivo and in vitro functional validation of its evolutionally conserved role in supporting HSPC expansion in zebrafish and mice. Finally, comparison between zebrafish CHT and human fetal liver highlighted the conservation and divergence across evolution.
Taken together, the researchers established a high-resolution single-cell transcriptomic atlas of HSPC expansion in the zebrafish CHT. These findings enhanced our understanding of the regulatory mechanism underlying hematopoietic niche for HSPC expansion in vivo and will provide insights into improving protocols for HSPC expansion in vitro.
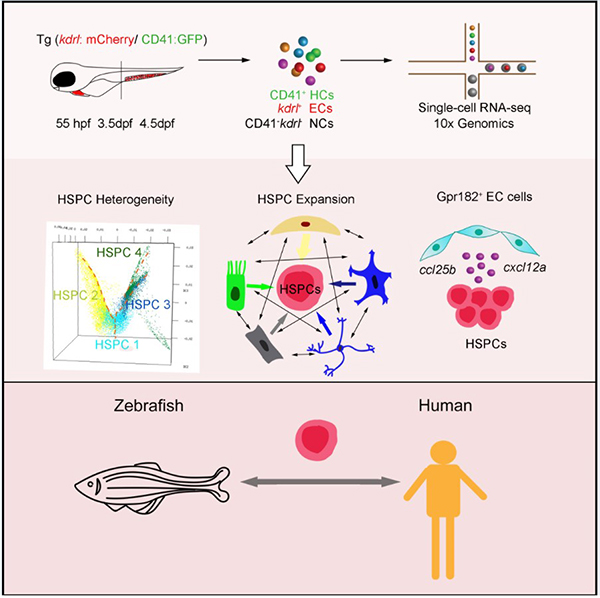

Figure 1. A single-cell resolution developmental atlas of hematopoietic stem and progenitor cell expansion in zebrafish. (Image by Prof. Feng Liu 's group)
Website: https://doi.org/10.1073/pnas.2015748118
1. E. Laurenti, B. Gottgens, From haematopoietic stem cells to complex differentiation landscapes. Nature 553, 418-426 (2018).
2. J. R. Perlin, A. L. Robertson, L. I. Zon, Efforts to enhance blood stem cell engraftment: Recent insights from zebrafish hematopoiesis. J Exp Med 214, 2817-2827 (2017).
3. C. R. Mantel et al., Enhancing Hematopoietic Stem Cell Transplantation Efficacy by Mitigating Oxygen Shock. Cell 161, 1553-1565 (2015).
4. A. C. Wilkinson, K. J. Igarashi, H. Nakauchi, Haematopoietic stem cell self-renewal in vivo and ex vivo. Nat Rev Genet 21, 541-554 (2020).
5. S. H. Orkin, L. I. Zon, Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631-644 (2008).
6. E. Murayama et al., Tracing hematopoietic precursor migration to successive hematopoietic organs during zebrafish development. Immunity 25, 963-975 (2006).
7. Y. Xue et al., The Vascular Niche Regulates Hematopoietic Stem and Progenitor Cell Lodgment and Expansion via klf6a-ccl25b. Dev Cell 42, 349-362 e344 (2017).
8. Y. Xue et al., A 3D Atlas of Hematopoietic Stem and Progenitor Cell Expansion by Multi-dimensional RNA-Seq Analysis. Cell Rep 27, 1567-1578 e1565 (2019).
(Contact: Prof. Feng Liu, liuf@ioz.ac.cn)


