The response of closely related species to shared environmental stress is likely to invoke congruent or similar physiological and/or morphological adaptations, a process termed “parallel evolution”. However, parallelism is typically stronger at the phenotypic level, while genetic solutions to achieve these phenotypic similarities may differ, especially for polygenic traits. Theoretical hypothesis suggests that such genetic nonparallelism may be influenced by the availability of standing genetic variation (i.e., heterozygosity). A loss of heterozygosity within small populations might reduce opportunities for parallel evolution across replicated species by decreasing the probability that the same alleles are available in response to shared selection pressures. Fumin Lei from the Institute of Zoology, Chinese Academy of Sciences and colleagues provided new eyes on this hypothesis in an article recently published in Proceedings of the National Academy of Sciences of the United States of America. Access to: https://doi.org/10.1073/pnas.2023918118.
To test the above hypothesis, they compared genomes of closely related, co-distributed tit species across East Asian continent. As East Asia is a region of extreme environmental variation. The eastern region of East Asia is at or near sea level, but western East Asia is dominated by the Qinghai–Tibet Plateau, with an average elevation of 4500 m a.s.l. During the Pleistocene, the western region was covered by glaciers, whereas the eastern lowlands were largely ice free. The tits and chickadees (Passeriformes: Paridae) that originated from the Sino-Himalayan region have 19 species distributing in both the eastern lowlands and western highlands throughout East Asia. Tits living in western highlands include western endemics and western populations of the widespread species (Fig. 1). Such species complexes consisting of species with variable demographic histories inhabiting extreme environments provide an opportunity to test this hypothesis.
Spatial estimation found that the heterozygosity was significantly lower in all western birds than the eastern ones (Fig. 2A). Specifically, western endemic species had consistently lower heterozygosity than eastern endemic species. However, the widespread species had a significantly higher heterozygosity than western endemics but similar to that observed in eastern endemics, although their western populations had a significantly lower heterozygosity than conspecific eastern populations (Fig. 2B). The inference of historical demography also indicated that all western endemics showed a significant decline in the effective population size during the Pleistocene (Fig. 2C). In contrast, most of eastern endemics showed significant demographic expansion during this period. For the widespread species, eastern populations showed significant population expansion, while western populations showed distinct demography from that in western endemics (Fig. 2D). This finding highlights the role of the demography in determining the level of heterozygosity.
To examine the extent to which high-elevation adaptation is parallel and test whether the level of parallelism is affected by heterozygosity among these high-elevation taxa with different heterozygosity levels and demography, this study further compared the most differentiated genomic regions between each high-elevation taxon and its low-elevation relative. Functional enrichment analyses found that the outlier regions in all 7 pairs were significantly enriched for genes potentially related to the oxygen transport cascade and/or thermogenesis (Fig. 3A). Despite no parallelism at particular genes, high similarity in gene function was found among comparisons, which focused on some functional categories like oxygen utilization, hypoxia responses, lipid metabolism and carbohydrate metabolism. Furthermore, parallelism is not higher in more heterozygous widespread parids than in western highland endemics (Fig. 3B). In conclusion, for East Asian parids, parallel response to extreme elevation appears to likely occur at functional or genomic levels relying on different genes, with differences in heterozygosity having no effect on the degree of genetic parallelism.
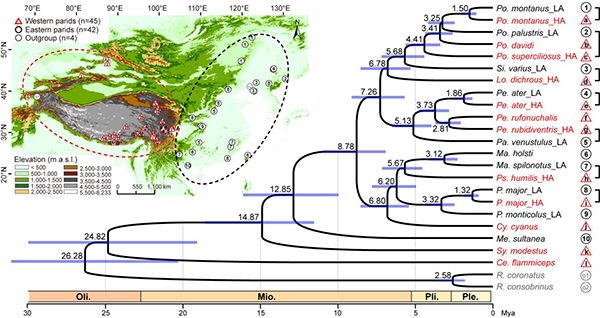
Fig. 1. Sampling localities and time tree of true tits in East Asia.
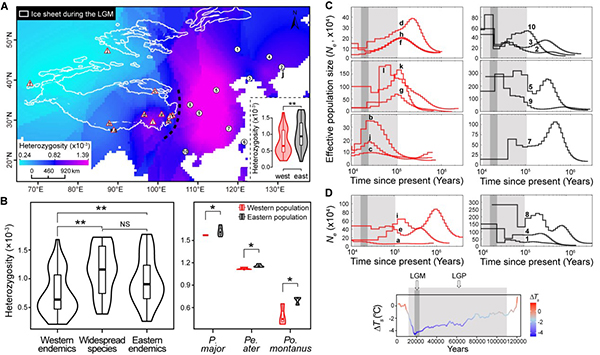
Fig. 2. (A) Spatial distribution of the heterozygosity. (B) Comparisons of heterozygosity levels among taxa. (C) Historical demography of western and eastern endemic tits. (D) Demography of widespread species.
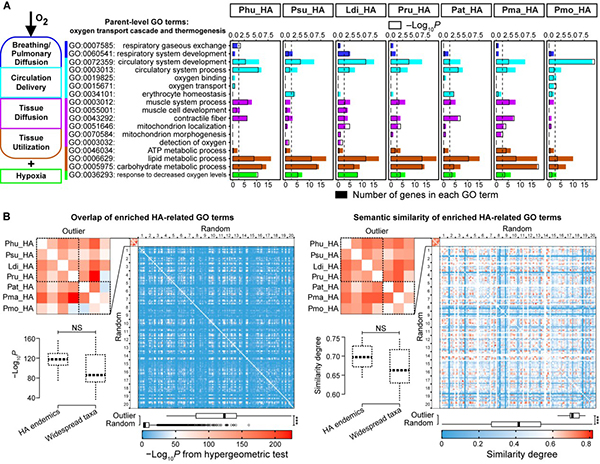
Fig. 3. (A) GO terms enriched by HA-related genes found in FST outlier windows. (B) Comparisons of similarity in HA-related GO terms.
(Contact:Fu-Min Lei, leifm@ioz.ac.cn)



